男女共同参画学協会連絡会
支援企業による広告記事
- ソーラボジャパン株式会社
- CAD を使って光学装置を設計してみよう
- 「生物物理」2025年10月号
- ソーラボジャパン株式会社
- 次世代2光子顕微鏡―小型化がもたらす新たな可能性
- 「生物物理」2024年10月号
- ソーラボジャパン株式会社
- サイエンティフィックカメラと周辺機器の同期
- 「生物物理」2023年10月号
- ソーラボジャパン株式会社
- 顕微鏡のリノベーション ~ 顕微鏡ポートを活用した光学系の導入
- 「生物物理」2022年12月号
「Biophysics and Physicobiology」に Seiichiro Hayashi, Daisuke Fujinami, Daisuke Kohda による "NMR evidence for residue-based LFER relationship in two-state folding-unfolding equilibrium of the spectrin SH3 domain in acidic solutions" をJ-STAGEの早期公開版として掲載
2026年03月05日 学会誌
日本生物物理学会欧文誌[Biophysics and Physicobiology]に以下の論文が早期公開されました。
Seiichiro Hayashi, Daisuke Fujinami, Daisuke Kohda
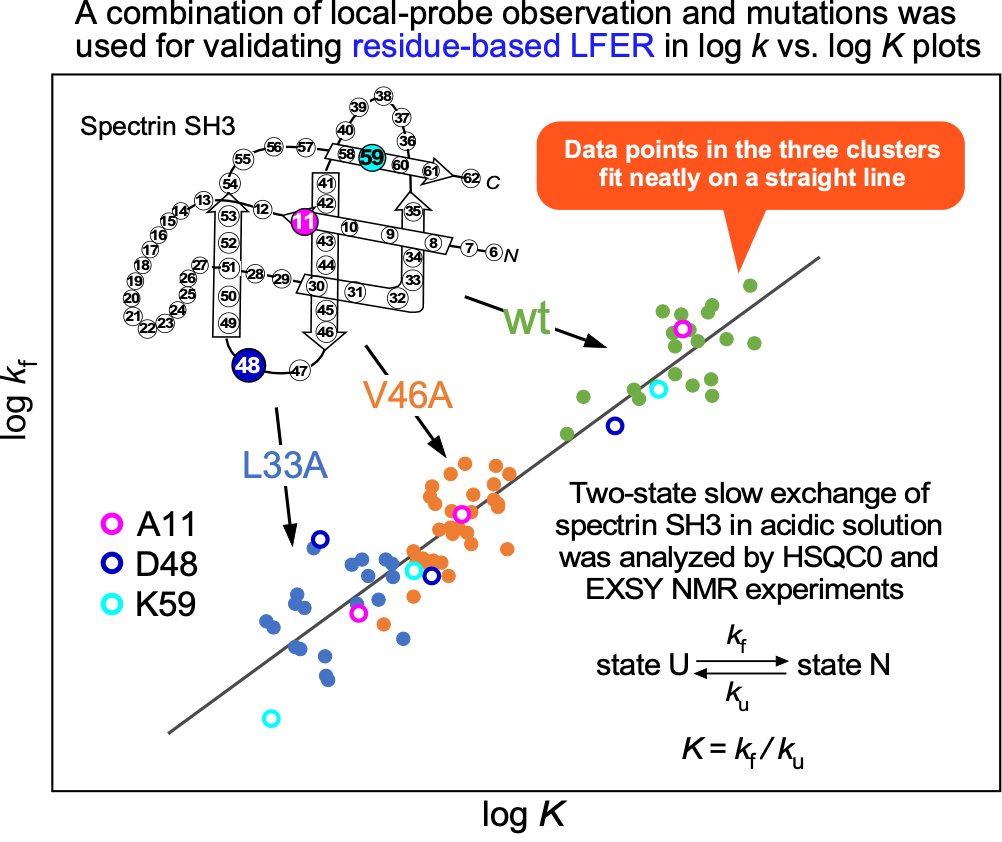
"NMR evidence for residue-based LFER relationship in two-state folding-unfolding equilibrium of the spectrin SH3 domain in acidic solutions"
URL:https://doi.org/10.2142/biophysico.bppb-v23.0011
- Abstract
- The conformational change of a protein molecule is described by the single equilibrium constant K and the single rate constants, k (forward) and k' (reverse). The underlying assumption is that all amino acid residues undergo state changes with perfect cooperativity. However, residue-specific measurements such as NMR often show residue-to-residue variations in these constants, indicating that proteins are not fully cooperative. Theoretical considerations based on “the consistency principle of protein folding” demonstrated that the linear free energy relationship (LFER) observed in residue-specific log k vs. log K plots is the physicochemical basis for smooth conformational changes of protein molecules. The residue-based LFER was found in many protein-related phenomena, but the structural changes have been limited to relatively small ones, such as fluctuations in the folded state and the coupled binding and folding of intrinsically disordered proteins. Here, we applied NMR to the two-state exchange equilibrium of the spectrin SH3 domain in acidic solutions and determined residue-specific equilibrium and rate constants. The data points obtained from the wild-type and its two single amino acid mutants were aligned on a single line in the log k vs. log K plot, indicating that residue-based LFER applies to large structural changes between the unfolded and folded states of the SH3 domain. The mutation-induced shifts in the distribution ranges of residue-specific equilibrium and rate constants are useful for establishing residue-based LFERs for all kinds of protein structural changes, including the refolding from a fully unfolded state, caused by the removal of a denaturant.
URL: https://doi.org/10.2142/biophysico.bppb-v23.0011